The RigidCLL library
RigidCLL Class ReferenceThe main RigidCLL class. More...
Collaboration diagram for RigidCLL:

Detailed DescriptionClass encompassing the information and methods implementing the RigidCLL algorithm. Definition at line 214 of file RigidCLL.h. Constructor & Destructor Documentation
The RigidCLL constructor. Definition at line 440 of file RigidCLL.cpp.
The RigidCLL constructor. Read information from a file to define a RigidCLL structure.
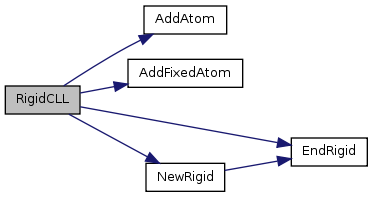
Definition at line 445 of file RigidCLL.cpp. References AddAtom(), AddFixedAtom(), EndRigid(), and NewRigid().
Here is the call graph for this function:
The RigidCLL destructor. Definition at line 527 of file RigidCLL.cpp. References Reset().
Here is the call graph for this function:
 Member Function Documentation
Returns the number of rigids in the RigidCLL structure. Note that the fixed atoms are counted as one rigid cluster. Thus, the output of this function is the number of mobile rigid groups plus one. If there is a rigid under construction, it is considered as compleately defined (altthogh some atoms can be defined latter on to it). Definition at line 493 of file RigidCLL.cpp. Referenced by main().
Here is the caller graph for this function:

Sets the cutoff. Use to to detect steric clashes (this is the default value). When using this function the internal grid is deleted (if any) and it will be re-defined the next call to an interaction detection method.
Definition at line 499 of file RigidCLL.cpp. References ERROR. Referenced by main().
Here is the caller graph for this function:
Returns the cutoff being used to detect atom interactions. This is basically the parameter given when calling SetCutOff. When aiming for steric clash detection, the returned cutoff is 0.
Definition at line 511 of file RigidCLL.cpp. Referenced by main().
Here is the caller graph for this function:

Size of the area coveed by the grid.
Definition at line 516 of file RigidCLL.cpp. References ERROR. Referenced by main().
Here is the caller graph for this function:
Defines a new fixed atom.
Definition at line 532 of file RigidCLL.cpp. References ERROR. Referenced by AddFixedAtom(), and RigidCLL().
Here is the caller graph for this function:
Defines a new fixed atom.
Definition at line 541 of file RigidCLL.cpp. References AddFixedAtom().
Here is the call graph for this function:

Starts the definition of a rigid group of atoms.
Definition at line 546 of file RigidCLL.cpp. References EndRigid(), ERROR, RigidInfo::firstAtom, and MAX_UINT. Referenced by RigidCLL().
Here is the call graph for this function:

Here is the caller graph for this function:

Defines a new atom in the current rigid group of atoms.
Definition at line 562 of file RigidCLL.cpp. References ERROR, RigidInfo::firstAtom, and MAX_UINT. Referenced by AddAtom(), and RigidCLL().
Here is the caller graph for this function:
Defines a new atom in the current rigid group of atoms.
Definition at line 571 of file RigidCLL.cpp. References AddAtom().
Here is the call graph for this function:

Concludes the definition of a rigid group of atoms. Definition at line 576 of file RigidCLL.cpp. References ERROR, RigidInfo::firstAtom, RigidInfo::lastAtom, and MAX_UINT. Referenced by NewRigid(), and RigidCLL().
Here is the caller graph for this function:

Activates the interaction between a pair of rigids. The interactions are set the first time we call ActivateInteraction or DeactivateInteraction. If not set, the interactions are assumed to be all active. When we start giving allowed or disabled intereactions, we can not add more rigids until we call the Reset method. Note that rigids are numbered from 1 in the order they are defined by the user. Rigid 0 refers to the 'fixed' group of atoms.
Definition at line 590 of file RigidCLL.cpp. References ERROR.
The interactions are set the first time we call ActivateInteraction or DeactivateInteraction. If not set, the interactions are assumed to be all active. When we start giving allowed or disabled intereactions, we can not add more rigids until we call the Reset method. Note that rigids are numbered from 1 in the order they are defined by the user. Rigid 0 refers to the 'fixed' group of atoms.
Definition at line 605 of file RigidCLL.cpp. References ERROR.
Intializes the structures required for collision detection. Once the WarmUp is used, no more rigids or fixed atoms can be added until we use the Reset method. In general there is no need to explicitly call this method, as, if necessary it will be automatically invoked when first checking for interactions.
Definition at line 616 of file RigidCLL.cpp. References BB_MARGIN, ERROR, INF, and MAX_UINT. Referenced by CLLInteractions(), CLLStericClash(), Interactions(), main(), and StericClash().
Here is the caller graph for this function:
Resets the internal information so that new rigids can be added to the problem. Note that both the grid and the interaction table will be deleted. Thus, the interaction information must be re-entered before checking for interaction (or it will be assumed that all rigid clusters can interact between them). Definition at line 704 of file RigidCLL.cpp. Referenced by ~RigidCLL().
Here is the caller graph for this function:

Determines if there is at least one steric clash in the problem. This is the method to be used when the WarmUp is called with cutoff 0.
Definition at line 710 of file RigidCLL.cpp. References ERROR, AtomInfo::move(), AtomInfo::rID, and WarmUp(). Referenced by main().
Here is the call graph for this function:
Here is the caller graph for this function:
StericClash using the orignal CLL algorithm (i.e., Algorithm 2 in the paper). This is used to show the advantage of the RigidCLL algorithm. Note that this methods takes into account the decomposition of the problem in rigid clusters and, thus, it only takes into account clashes between atoms in different clusters. This is the same as StericClash.
Definition at line 763 of file RigidCLL.cpp. References ERROR, and WarmUp(). Referenced by main().
Here is the call graph for this function:

Here is the caller graph for this function:

A brute force alternative to StericClash used for test and debug.
Definition at line 801 of file RigidCLL.cpp. References ERROR, MANHATTAN_DISTANCE, AtomInfo::r, and AtomInfo::rID. Referenced by main().
Here is the caller graph for this function:

Detects nearby atoms (atoms closer than the cutoff). This is the method to sue when the WarmUp is called with cutoff larger than 0. Note that only pair of atoms in different rigid groups are considered. This is the essence of the RigidCLL algorihtm.
Definition at line 857 of file RigidCLL.cpp. References AtomInfo::c, ERROR, RigidInfo::firstAtom, CellInfo::fixedID, MANHATTAN_DISTANCE, MAX_UINT, CellInfo::mobileID, AtomInfo::rID, and WarmUp(). Referenced by main().
Here is the call graph for this function:

Here is the caller graph for this function:

The same as Interactions but using the standard CLL algoritm. This is used to show the advantage of the RigidCLL algorithm. Note that this methods does not take into account the decomposition of the problem in rigid clusters and, thus, it can return interactions between atoms in the same rigid (in contrast, Interactions only considers interactions between atoms in different clusters). Thus, the list of interactions returned by this method is typically larger than the one returned by Interactions
Definition at line 958 of file RigidCLL.cpp. References AtomInfo::c, ERROR, CellInfo::fixedID, AtomInfo::id, MANHATTAN_DISTANCE, MAX_UINT, CellInfo::mobileID, and WarmUp(). Referenced by main().
Here is the call graph for this function:

Here is the caller graph for this function:

A brute force alternative to Interactions used for test and debug. This method follows the same policy as Interactions (and not the one of CLLInteractions): it only returns interactions between atoms in different rigid clusters. Thus, the output of this method can be used to check the output of Interactions but not the one of CLLInteractions.
Definition at line 1043 of file RigidCLL.cpp. References ERROR, MANHATTAN_DISTANCE, and AtomInfo::rID. Referenced by main().
Here is the caller graph for this function:
 The documentation for this class was generated from the following files: | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||

Follow us!