|
parameters.h File Reference Detailed DescriptionDefinition of the Tparameters type and the associated functions.
Definition in file parameters.h.
Macro Definition Documentation
In this representation we use 6 parameters to represent two vectors of the rotation matrices for each link. The third vector is the cross product of the two represented vectors.
Definition at line 27 of file parameters.h. Referenced by ApplyLinkRot(), CacheRotVars(), GenerateLinkRot(), GenerateLinkSolution(), GetLinkPoseSimpVars(), GetTransform2Link(), PrintParameters(), RegenerateLinkBox(), RegenerateLinkSolution(), and SetRotVars().
In this representation we use 9 parameters to represent the full rotation matrices for each link.
Definition at line 37 of file parameters.h. Referenced by ApplyLinkRot(), CacheRotVars(), GenerateLinkRot(), GenerateLinkSolution(), GetLinkPoseSimpVars(), GetTransform2Link(), PrintParameters(), RegenerateLinkBox(), RegenerateLinkSolution(), and SetRotVars().
In this representation we use 4 parameters to represent rotation matrices for each link relying on quaternions.
Definition at line 48 of file parameters.h. Referenced by ApplyLinkRot(), CacheRotVars(), ChangeLinkReferenceFrame(), GenerateLinkRot(), GenerateLinkSolution(), GetLinkPoseSimpVars(), GetTransform2Link(), PrintParameters(), RegenerateLinkBox(), RegenerateLinkSolution(), and SetRotVars().
In this representation we do not use any variable for links but only for joints. This is similar to the classical DH non redundant formulation, but includes some auxliary/dummy variables for some kind of joints (in_patch).
Definition at line 60 of file parameters.h. Referenced by ApplyLinkRot(), ApplyLinkRotVar(), CacheRotVars(), ChangeLinkReferenceFrame(), GenerateEquationsFromBranch(), GenerateJointEquations(), GenerateJointEquationsInBranch(), GenerateJointRangeEquations(), GenerateJointRangeSingularityEquations(), GenerateJointSolution(), GenerateLinkRot(), GenerateLinkSolution(), GenerateTransEquationsFromBranch(), GenerateWorldEquations(), GenerateWorldSingularityEquations(), GetJointTransSeq(), GetLinkPoseSimpVars(), GetLinkTransformsFromSolution(), GetLinkTransformsFromSolutionPoint(), GetSolutionPointFromLinkTransforms(), GetTransform2Link(), InitWorldKinCS(), main(), MoveWorld(), NewRevoluteJoint(), NewUniversalJoint(), PlotChartAsPolygon(), PrintParameters(), PrintWorldAxes(), ReadOneSample(), ReadTwoSamples(), RegenerateJointBox(), RegenerateJointSolution(), RegenerateLinkBox(), RegenerateLinkSolution(), RegenerateMechanismBox(), RegenerateWorldOriginalPoint(), SetRotVars(), TriangulateAtlas(), WorldAtomJacobian(), WorldDOF2Sol(), and WorldSample2DOF().
One of the possible collison detection engines. Definition at line 69 of file parameters.h. Referenced by CheckCollision(), DeleteCD(), InitCD(), PrintCollisionInfo(), PrintParameters(), and StoreCollisionInfo().
One of the possible collison detection engines. Definition at line 78 of file parameters.h. Referenced by CheckCollision(), DeleteCD(), InitCD(), and PrintParameters().
One of the possible collison detection engines. Definition at line 87 of file parameters.h. Referenced by CheckCollision(), DeleteCD(), InitCD(), and PrintParameters().
One of the possible collison detection engines. Definition at line 96 of file parameters.h. Referenced by CheckCollision(), DeleteCD(), InitCD(), PrintCollisionInfo(), PrintParameters(), and StoreCollisionInfo().
One of the possible collison detection engines. Definition at line 105 of file parameters.h. Referenced by CheckCollision(), DeleteCD(), InitCD(), MoveAndCheckCD(), PrintParameters(), and WorldContinuousCD().
One of the possible collison detection engines. Definition at line 114 of file parameters.h. Referenced by CheckCollision(), DeleteCD(), InitCD(), PrintCollisionInfo(), and StoreCollisionInfo().
One of the possible collison detection engines. Definition at line 123 of file parameters.h. Referenced by CheckCollision(), DeleteCD(), and InitCD().
One of the possible collison detection engines. Definition at line 132 of file parameters.h. Referenced by CheckCollision(), DeleteCD(), InitCD(), and PrintParameters().
One of the possible collison detection engines. Definition at line 141 of file parameters.h. Referenced by CheckCollision(), DeleteCD(), InitCD(), and PrintParameters().
In this sampling mode, points are generated randomly in the ambient space box.
Definition at line 151 of file parameters.h. Referenced by AtlasRRTSample(), and PrintParameters().
In this sampling mode, points are generated randomly in the boxes bounding the tree nodes. These boxes are defined in the kd-tree.
Definition at line 162 of file parameters.h. Referenced by AtlasRRTSample(), InitRRT(), PrintParameters(), and RRTSample().
In this sampling mode, points are generated randomly in the tangent spaces stored in the charts. This is only used in AtlasRRT. If the sampling mode is set to this value normal RRTs use AMBIENT_SAMPLING
Definition at line 174 of file parameters.h. Referenced by AtlasRRT(), AtlasRRTSample(), PointTowardRandSample(), and PrintParameters().
Default file of parameters. This is the file of parameters read in the first place, before reading the file specific for the problem at hand. In this way, the problem specific parameter need only to include values for the parameters whose value is not to be the default value. Definition at line 185 of file parameters.h. Referenced by InitParametersFromFile().
Threshold to turn any number into 0 before passing it to the simplex. This should be larger than 1e-6, the threshold hard-coded in many simplex packages. Definition at line 194 of file parameters.h. Referenced by AddBranchToAtlasRRT(), AddNodeToRRT(), AddStepToAtlasRRTstar(), AddStepToRRTstar(), AtlasAStar(), AtlasGBF(), AtlasRRT(), AtlasRRTstar(), AtlasTRRT(), cBiRRT(), ccRRT(), ccTRRT(), Chart2Manifold(), ClassifyPointInChart(), ComputeSplitDimInt(), ConnectSamples(), ConnectSamplesChart(), CSRemoveLCVars(), CSRemoveUnusedVars(), CSRemoveVarsWithCtRange(), CuikNewtonInBox(), CuikNewtonSimp(), ExtendAtlasFromPoint(), ExtendAtlasTowardPoint(), FindSingularPoint(), GetChartDegree(), IncrementalSampleCuikSystemInBox(), InitAtlasFromPoint(), InitChartInt(), InitRRT(), main(), MinimizeOnAtlas(), NewChartFromPoint(), NewTemptativeSample(), Newton2ManifoldPlane(), PointOnChart(), PopulateWithSamples(), PostProcessBox(), RecursiveReWireRRTstar(), ReduceBox(), ReduceBoxEquationWise(), RefineSingularPoint(), RegenerateSolution(), ReWireAtlasRRTstar(), ReWireRRTstar(), RRTstar(), SampleCuikSystemInBox(), Steps2PathinAtlasRRT(), Steps2PathinRRT(), TransitionTestRRT(), UpdateCuikSystem(), WireAtlasRRTstar(), and WireRRTstar().
The mechanisms configurations can be represented in different ways
Note that if you change this parameter solutions files previosly computed can become invalid. This can be a source of confusion so care must be taken to create/manipulate solutions using the same value for this parameter. Definition at line 215 of file parameters.h. Referenced by ApplyLinkRot(), ApplyLinkRotVar(), GenerateEquationsFromBranch(), GenerateJointEquations(), GenerateJointEquationsInBranch(), GenerateJointRangeEquations(), GenerateJointRangeSingularityEquations(), GenerateJointSolution(), GenerateLinkRot(), GenerateLinkSolution(), GenerateTransEquationsFromBranch(), GenerateWorldEquations(), GenerateWorldSingularityEquations(), GetJointTransSeq(), GetLinkPoseSimpVars(), GetLinkTransformsFromSolution(), GetLinkTransformsFromSolutionPoint(), GetSolutionPointFromLinkTransforms(), GetTransform2Link(), InitWorldFromMolecule(), InitWorldKinCS(), main(), MoveWorld(), PlotChartAsPolygon(), PrintParameters(), PrintWorldAxes(), ReadOneSample(), ReadTwoSamples(), RegenerateJointBox(), RegenerateJointSolution(), RegenerateLinkBox(), RegenerateLinkSolution(), RegenerateMechanismBox(), RegenerateWorldOriginalPoint(), SetRotVars(), TriangulateAtlas(), WorldAtomJacobian(), WorldDOF2Sol(), and WorldSample2DOF().
If boxes reduce more that RHO the reduction process is repeated. Definition at line 222 of file parameters.h. Referenced by ReduceBox(), ReduceBoxEquationWise(), and RegenerateSolution().
Size of the boxes bounding isolated solutions. Definition at line 229 of file parameters.h. Referenced by IncrementalSampleCuikSystemInBox(), ReduceBox(), ReduceBoxEquationWise(), and SampleCuikSystemInBox().
Size of the boxes bounding continuous solution sets. Definition at line 236 of file parameters.h. Referenced by PostProcessBox().
Maximum projection error of the local maps when building atlas (or part of atlas). Definition at line 243 of file parameters.h. Referenced by ConnectSamplesChart(), GradientSmooth(), InitAtlasFromPoint(), InitAtlasRRT(), and main().
Maximum oriented curvature error of the local maps when building atlas (or part of atlas). Definition at line 250 of file parameters.h. Referenced by ConnectSamplesChart(), GradientSmooth(), InitAtlasFromPoint(), InitAtlasRRT(), and main().
Maximum radius of the local maps when building atlas (or part of atlas). Definition at line 257 of file parameters.h. Referenced by ConnectSamplesChart(), GradientSmooth(), InitAtlasFromPoint(), InitAtlasRRT(), main(), and SPolytope2Polytope().
Radius of the ball used to generate samples. This must be larger than CT_R.
Definition at line 266 of file parameters.h. Referenced by InitChartInt(), and Polytope2SPolytope().
Step size for the paths connecting two given configurations. In between two steps no collision detetion is performed so, indirectly, this parameter defines the maximum penetrability between any two bodies in the problem. Note that points in the path connecting two configurations are determined via linear interpolation between the start and goal configurations and then projecting on the manifold. The projection step (different for different algorithms) slightly changes the distance between consecutive steps in the path. Therefore, this parameter is only an approximation of the final actual value. Definition at line 282 of file parameters.h. Referenced by AddBranchToAtlasRRT(), AtlasBiRRTstar(), AtlasRRTstar(), ConnectSamples(), ConnectSamplesChart(), DistanceOnChart(), ExtendAtlasTowardPoint(), GeodesicDistance(), GradientSmooth(), InitChartInt(), InitRRT(), MinimizeOnAtlas(), PathInChart(), Polytope2SPolytope(), and SmoothSamples().
Cost penalty factor for atlasGBF. Definition at line 289 of file parameters.h. Referenced by AtlasAStar(), AtlasGBF(), and BuildAtlasFromPoint().
Period between two consecutive savings of the internal cuik state. Definition at line 296 of file parameters.h. Referenced by MPI_SolveCuikSystem(), and SolveCuikSystem().
Number of output solutions boxes we need. Use 0 to obtain all possible solution boxes for the problem at hand. Definition at line 304 of file parameters.h. Referenced by DealWithCP(), IncrementalSampleCuikSystemInBox(), MPI_SolveCuikSystem(), SampleCuikSystemInBox(), and SolveCuikSystem().
Maximum number iterations in the Newton-Raphson function. Definition at line 311 of file parameters.h. Referenced by Chart2Manifold(), CuikNewtonInBox(), CuikNewtonSimp(), IncrementalSampleCuikSystemInBox(), main(), Newton2ManifoldPlane(), PostProcessBox(), and RefineSingularPoint().
Dimensionality of the solution space for the mechanism at hand. Definition at line 318 of file parameters.h. Referenced by BuildAtlasFromPoint(), ConnectSamplesChart(), DealWithCP(), GetSCpSystem(), GradientSmooth(), InitAtlasFromPoint(), InitAtlasRRT(), InitRRT(), main(), MinimizeOnAtlas(), RandomSmooth(), and ShortcutSmooth().
Contant part of the search radius for nearest neightours in RRT*. Definition at line 325 of file parameters.h. Referenced by AtlasBiRRTstar(), AtlasRRTstar(), BiRRTstar(), and RRTstar().
Dummification level.
Definition at line 339 of file parameters.h. Referenced by DummifyAndAddEquation(), and main().
TRUE to use a linearization error minimization criterion to select the split dimension. FALSE to split along the largest side. Definition at line 346 of file parameters.h. Referenced by ComputeSplitDimInt(), PrintParameters(), and SampleCuikSystemInBox().
Trade off between speed and numerical stability when using the simplex Possible values are
Definition at line 358 of file parameters.h. Referenced by ReduceBox().
Simplification level:
The less the simplification the more the numerical stability, but the slower the solution process. A simplification level 2 should be Ok except for extremely ill conditioned cases. Definition at line 378 of file parameters.h. Referenced by PrintCuikSystemWithSimplification(), and SimplifyCuikSystem().
Threshold to switch from a specific linearization (one based on linear relaxations) to a general one (based on Taylorm models) for purely quadratic equations: parabolas, circles, and spheres. Definition at line 389 of file parameters.h. Referenced by ReduceBox().
Threshold to switch from a specific linearization (one based on linear relaxations ) to a general one (based on Taylorm models) for saddle equations. Definition at line 398 of file parameters.h. Referenced by ReduceBox().
Reduction ratio [0,1] over the Van der Waals radius used to detect steric clashes. The reduction is used to ensure that only very high energy conformations are discarded by considering the collisions between atoms (as if atoms where solid spheres). Definition at line 408 of file parameters.h. Referenced by InitWorldFromMolecule().
Limit of domain of the X dimension of 3d plots when the variables have circular ranges. The default limits are [-pi,pi] and this sometimes produce cutted plots. Using these constants the range of the plot can be adjusted at will to avoid those cuts. This is basically used when plotting atlas the DOF CT_REPRESENTATION. Definition at line 420 of file parameters.h. Referenced by PlotChart(), PlotChartAsPolygon(), PlotRRT(), PlotSamples(), PrintParameters(), and TriangulateAtlas().
Limit of domain of the Y dimension of 3d plots when the variables have circular ranges. The default limits are [-pi,pi] and this sometimes produce cutted plots. Using these constants the range of the plot can be adjusted at will to avoid those cuts. This is basically used when plotting atlas the DOF CT_REPRESENTATION. Definition at line 432 of file parameters.h. Referenced by PlotChart(), PlotChartAsPolygon(), PlotRRT(), PlotSamples(), PrintParameters(), and TriangulateAtlas().
Limit of domain of the Z dimension of 3d plots when the variables have circular ranges. The default limits are [-pi,pi] and this sometimes produce cutted plots. Using these constants the range of the plot can be adjusted at will to avoid those cuts. This is basically used when plotting atlas the DOF CT_REPRESENTATION. Definition at line 444 of file parameters.h. Referenced by PlotChart(), PlotChartAsPolygon(), PlotRRT(), PlotSamples(), PrintParameters(), and TriangulateAtlas().
Scale factor of the temperature parameter for Transition-based RRT Definition at line 451 of file parameters.h. Referenced by TransitionTestRRT().
Number of successive trnasition test failues requiered before increasing the temperature Definition at line 460 of file parameters.h. Referenced by TransitionTestRRT().
To activate/deactivate the search for bifurcations during the atlas construction. Definition at line 468 of file parameters.h. Referenced by AddTrustedChart2Atlas(), AtlasRRT(), AtlasTRRT(), DetermineChartNeighbours(), main(), MinimizeOnAtlas(), and PrintAtlasStatistics().
Maximum time in seconds for the path planning algorihtms (cuikatlasGBF, cuikaltasrrt, cuikccrrt,....). Definition at line 475 of file parameters.h. Referenced by AtlasAStar(), AtlasBiRRTstar(), AtlasGBF(), AtlasRRT(), AtlasRRTstar(), AtlasTRRT(), BiRRTstar(), cBiRRT(), ccRRT(), ccTRRT(), and RRTstar().
Maximum iterations for the path planning algorihtms (currently only used in AtlasRRT* and RR* since the rest of algorithms are limited in time only). Definition at line 483 of file parameters.h. Referenced by AtlasBiRRTstar(), AtlasRRTstar(), BiRRTstar(), main(), and RRTstar().
Maximum number of charts in at atlas. Definition at line 490 of file parameters.h. Referenced by AtlasAStar(), AtlasGBF(), and BuildAtlasFromPoint().
Maximum number of nodes in an RRT. Definition at line 497 of file parameters.h. Referenced by AddBranchToAtlasRRT(), AtlasRRT(), AtlasTRRT(), ccRRT(), and ccTRRT().
TRUE if the RRT search is bi-directional. cuikccrrt is one-directional despite the value of this parameter. Definition at line 508 of file parameters.h. Referenced by main().
TRUE if the RRT has to store all the neighbours for each new node. This is only used in cuikatlasrrtstar to speed up the propagation of improvements of the path to the nodes. Definition at line 518 of file parameters.h. Referenced by main().
TRUE if the RRT-like planners have to use dynamic domains. See the paper:
for more details on this technique. Definition at line 533 of file parameters.h. Referenced by InitRRT().
Collision detection engine. Right now four options are available: SOLID, VCOLLIDE, PQP, FCL, C_FCL, BULLET,RigidCD, NONE. Definition at line 542 of file parameters.h. Referenced by InitWorldCD(), and PrintParameters().
Parameter used to select the mode to generate samples to expand the (Atlas)RRT. Three modes are available: TANGENT_SAMPLING, AMBIENT_SAMPLING, KDTREE_SAMPLING. TANGENT_SAMPLING is the default mode for Atlas RRT and uses the charts to sample. AMBIENT_SAMPLING is the default mode for RRTs. This mode generate samples from the ambient space. KDTREE_SAMPLING is a mode where the samples are generated from boxes bounding the nodes in the RRT. This boxes are defined by a kd-tree (also used to accelerate the searches for neighbour nodes). Definition at line 563 of file parameters.h. Referenced by AddBranchToAtlasRRT(), AtlasBiRRTstar(), AtlasRRT(), AtlasRRTstar(), AtlasTRRT(), BiRRTstar(), cBiRRT(), ccRRT(), ccTRRT(), InitRRT(), NewTemptativeSample(), PrintParameters(), and RRTstar().
Total number of parameters. Definition at line 570 of file parameters.h. Referenced by ChangeParameter(), GetParameter(), GetParameterID(), InitParameters(), InitParametersFromFile(), PrintParameters(), and SetParameter(). Typedef Documentation
A set of parameters. Each parameter is identified with a constant defined in file parameters.h. This is basically a set of constants, but only a small, predefined set of constants are allowed. This is why we do not re-use Tconstants but provide a different implementation.
Definition at line 606 of file parameters.h. Function Documentation
Defines an empty set of parameters.
Definition at line 36 of file parameters.c. References NPARAMETERS. Referenced by InitParametersFromFile().
Defines an set of parameters from a file. It first read the default set of parameters and then the given set of parameters.
Definition at line 51 of file parameters.c. References CreateFileName(), DEFAULT_PARAMS, DeleteFileName(), Error(), GetFileFullName(), InitParameters(), NPARAMETERS, PARAM_EXT, ReadParameters(), and Warning(). Referenced by main().
Gets the value for a particular parameter.
Definition at line 93 of file parameters.c. References Error(), and NPARAMETERS. Referenced by AddBranchToAtlasRRT(), AddNodeToRRT(), AddStepToAtlasRRTstar(), AddStepToRRTstar(), AddTrustedChart2Atlas(), ApplyLinkRot(), ApplyLinkRotVar(), AtlasAStar(), AtlasBiRRTstar(), AtlasGBF(), AtlasRRT(), AtlasRRTstar(), AtlasTRRT(), BiRRTstar(), BuildAtlasFromPoint(), cBiRRT(), ccRRT(), ccTRRT(), Chart2Manifold(), ClassifyPointInChart(), ComputeSplitDimInt(), ConnectSamples(), ConnectSamplesChart(), CSRemoveLCVars(), CSRemoveUnusedVars(), CSRemoveVarsWithCtRange(), CuikNewtonInBox(), CuikNewtonSimp(), DealWithCP(), DetermineChartNeighbours(), DistanceOnChart(), DummifyAndAddEquation(), ExtendAtlasFromPoint(), ExtendAtlasTowardPoint(), FindSingularPoint(), GenerateEquationsFromBranch(), GenerateJointEquations(), GenerateJointEquationsInBranch(), GenerateJointRangeEquations(), GenerateJointRangeSingularityEquations(), GenerateJointSolution(), GenerateLinkRot(), GenerateLinkSolution(), GenerateTransEquationsFromBranch(), GenerateWorldEquations(), GenerateWorldSingularityEquations(), GeodesicDistance(), GetChartDegree(), GetJointTransSeq(), GetLinkPoseSimpVars(), GetLinkTransformsFromSolution(), GetLinkTransformsFromSolutionPoint(), GetSCpSystem(), GetSolutionPointFromLinkTransforms(), GetTransform2Link(), GradientSmooth(), IncrementalSampleCuikSystemInBox(), InitAtlasFromPoint(), InitAtlasRRT(), InitChartInt(), InitRRT(), InitWorldCD(), InitWorldFromMolecule(), InitWorldKinCS(), main(), MinimizeOnAtlas(), MoveWorld(), MPI_SolveCuikSystem(), NewChartFromPoint(), NewTemptativeSample(), Newton2ManifoldPlane(), PathInChart(), PlotChart(), PlotChartAsPolygon(), PlotRRT(), PlotSamples(), PointOnChart(), Polytope2SPolytope(), PopulateWithSamples(), PostProcessBox(), PrintAtlasStatistics(), PrintCuikSystemWithSimplification(), PrintWorldAxes(), RandomSmooth(), ReadOneSample(), ReadTwoSamples(), RecursiveReWireRRTstar(), ReduceBox(), ReduceBoxEquationWise(), RefineSingularPoint(), RegenerateJointBox(), RegenerateJointSolution(), RegenerateLinkBox(), RegenerateLinkSolution(), RegenerateMechanismBox(), RegenerateSolution(), RegenerateWorldOriginalPoint(), ReWireAtlasRRTstar(), ReWireRRTstar(), RRTstar(), SampleCuikSystemInBox(), SetRotVars(), ShortcutSmooth(), SimplifyCuikSystem(), SmoothSamples(), SolveCuikSystem(), SPolytope2Polytope(), Steps2PathinAtlasRRT(), Steps2PathinRRT(), TransitionTestRRT(), TriangulateAtlas(), UpdateCuikSystem(), WireAtlasRRTstar(), WireRRTstar(), WorldAtomJacobian(), WorldDOF2Sol(), and WorldSample2DOF().
Determines the parameter identifier for a given parameter name.
Definition at line 110 of file parameters.c. References FALSE, NO_UINT, and NPARAMETERS.
Determines is a parameter is already defined. The default initialization (the one from the default set of parameters) is not taken into account. Only explicit initializations from the users are considered. Only parameters alrady defined can be used in the definition of other parameters.
Definition at line 131 of file parameters.c.
Sets the name and value for a particular parameter.
Definition at line 139 of file parameters.c. References ChangeParameter(), Error(), NPARAMETERS, and Warning().
Sets the value for a particular parameter.
Definition at line 164 of file parameters.c. References Error(), and NPARAMETERS. Referenced by DealWithCP(), IncrementalSampleCuikSystemInBox(), InitAtlasFromPoint(), main(), MinimizeOnAtlas(), SampleCuikSystemInBox(), and SetParameter().
Writes a parameter set to a stream, that can be stdout. In principle the output of this function can be used as a parameter file (i.e., it can be parsed correctly with InitParametersFromFile).
Definition at line 181 of file parameters.c. References AMBIENT_SAMPLING, C_FCL, CT_CD_ENGINE, CT_CUT_X, CT_CUT_Y, CT_CUT_Z, CT_REPRESENTATION, CT_SAMPLING, CT_SPLIT_TYPE, Error(), FCL, INF, KDTREE_SAMPLING, NOCD, NPARAMETERS, PQP, REP_FLINKS, REP_JOINTS, REP_LINKS, REP_QLINKS, RIGIDCLL, SOLID, TANGENT_SAMPLING, and VCOLLIDE. Referenced by main().
Deletes a set of parameters and frees the allocated memory.
Definition at line 295 of file parameters.c. Referenced by main(). | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
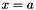 or
or 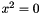 are considered (with x a variable and a a constant).
are considered (with x a variable and a a constant).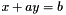 are also considered (with x and y variables and a and b constants).
are also considered (with x and y variables and a and b constants).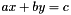 are also considered (with x and y variables and a and b constants).
are also considered (with x and y variables and a and b constants).
Follow us!